In a matter of months, the COVID-19 pandemic has besieged humanity and now the world wrestles to manage the population health challenges of a novel coronavirus with remarkable infectivity.
Organizing an effective response to blunt the impact of such a large, complex challenge demands a principled and scientific approach.
Better Planning by Forecasting Infections
Reliable forecasting is crucial for planning and allocating limited resources efficiently and minimizing casualties.
A most important characteristic of an infective virus is its average rate of reproduction or $R_0$. If generally, the number of people infected by a single person is greater than one, the virus will experience a phase of exponential growth in the rate of transmission among a population of susceptible individuals.
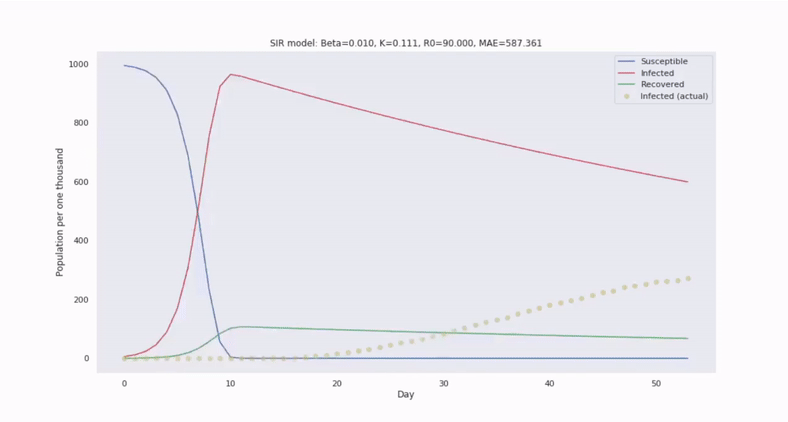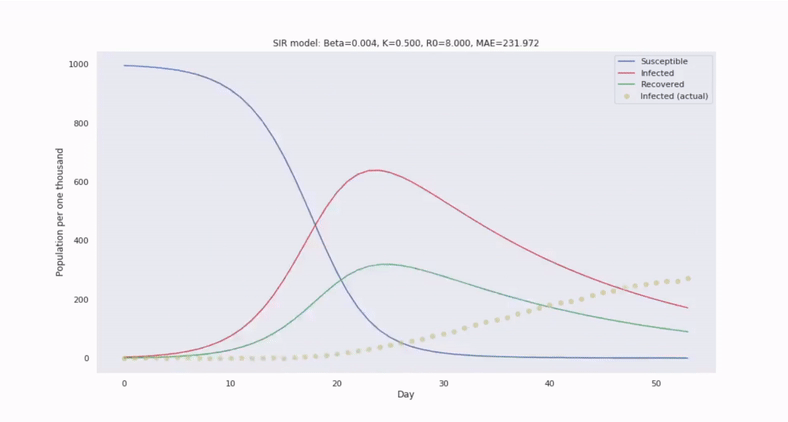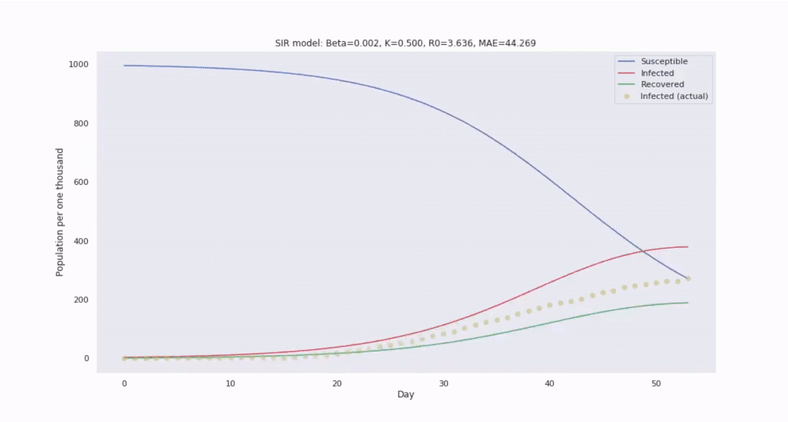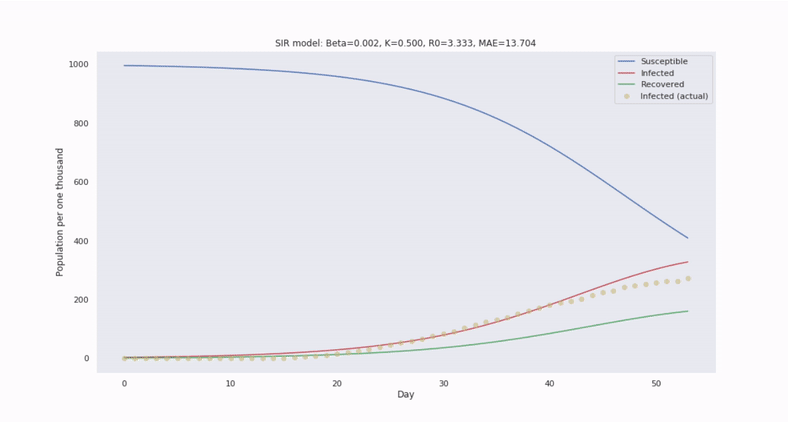
The canonical epidemiological model is called SIR and it segments a population into three disjoint subgroups: Susceptible, Infected, and Recovered (unable to become reinfected). SIR utilizes a system of nonlinear differential equations to describe how relative frequencies between these groups evolve in time.
Then to forecast the number of infected people in the future, we only need to solve the ODEs by integrating in time.
With SIR, we also assume the population is well-mixed. So, we can characterize the transition of susceptible to infected persons at the population level through the average contact rate of transmission, denoted by the parameter $\beta$, which scales an interaction cross term between $s(t)$ and $i(t)$.
Then $\beta s(t)i(t)$ individuals are removed from $S$ and added to $I$.
Similarly, the average rate of recovery is modeled by the parameter $\nu$ and $\nu i(t)$ individuals leave $I$ and enter $R$.
This yields the system:
\begin{equation} \frac{ds}{dt} = -\beta si \end{equation}
\begin{equation} \frac{di}{dt} = \beta si - \nu i \end{equation}
\begin{equation} \frac{dr}{dt} = \nu i \end{equation}
We can numerically approximate the derivatives using Euler’s method to produce the following update rules:
s_n1 = s_n - beta * s_n * i_n * del_t
i_n1 = i_n + (beta * s_n * i_n - k * i_n) * del_t
r_n1 = k * i_n
For small del_t, we can compute the variables S, I, and R at time n+1 in terms of known values at time n.
From these simple rules, we see an epidemic outbreak when $\frac{di}{dt} > 0$ which holds when: $\beta si - \nu i > 0$. At the outset, $S$ is is near 1 and so this holds for $\frac{\beta}{\nu} > 1$. The term $\frac{\beta}{\nu}$ is the same as $R_0$.
From this single constant, we can:
- Determine the rate of initial growth and final size of the epidemic
- Observe the effect of mitigation strategies like quarantine and vaccination
We can perform some qualitative analysis using the Andersen and May parametrization to reveal that the dynamics of this system are characterized by exponential growth for small time $t$.
Compared to regression models, we can forecast with greater confidence, despite limited and noisy data. This is because the SIR system of equations introduces greater constraint on the solution space.
Model parameters can be estimated by comparing with historical epidemics and fitting to observations. By simulating trajectories generated from a range of values, we can estimate the variability in the trajectories.
Groups like those at the University of Washington developed models based upon a related variant called SEIR while incorporating additional modeling techniques to make high quality short term predictions using additional sources such as geographic data.
Here you can read more about other variants which incorporate geographic structures or interactions between age groups with different prevalence rates.
The COVID-19 pandemic has also revealed a racial and socioeconomic bias in outcomes. These model shortcomings must be rectified to make ethically sound decisions.
We have seen regional healthcare infrastructures stressed to the brink of collapse under explosive outbreaks. But these geographically scattered events are also staggered in time, a fact we can use to avoid overwhelming the healthcare system.
Applications Optimizing Logistics
With the ability to reliably forecast infections, we can more efficiently use limited treatment resources like hospital beds, ventilators, and protective equipment.
Regarding healthcare providers as nodes in a distribution network, we can frame a transshipment distribution problem, which can be handled efficiently with solvers in or-tools.
Then we can efficiently surge scarce resources to the places they are most acutely needed to reduce strain on the system with this generalization of the minimum cost flow problem.
With some simplifying assumptions about the time and cost to transport resources, we can explore logistics scheduling based on meeting demand through redeploying underutilized assets.
From this principled baseline, we can incorporate additional context to make better decisions grounded in the sciences of epidemiological modeling and logistics.
Mining COVID-19 Research using Elasticsearch
One of our early applications used Elasticsearch to help clinicians review the medical history of members for prior authorization workflows in a large healthcare payor.
Clinical decision making was bottlenecked by the fact that member information was spread across different legacy systems. Much of this information was kept as .tif attachments from faxed documents coming in from the regional network of independent physicians.
To facilitate evidence-based decision making, we developed an OCR pipeline using Tesseract and indexed these documents along with member info from the organization’s various relational databases. The resulting application offered a snappy, integrated view of the clinical history for each member.
We improved the search results by indexing similar terms as measured by cosine similarity of word2vec embeddings as well as a knowledge graph constructed using the UMLS medical ontology. Then at query time, we expand the search to include synonyms retrieved from an secondary index.
The recent COVID-19 pandemic has framed the need to scale up research around the virus as well as public health mitigation strategies & treatment methodologies.
Kaggle is hosting a challenge to develop information retrieval tools to help researchers mine the emerging corpus of literature.
Both NLP and Elasticsearch have evolved considerably since our work in building a clinical search tool to support prior authorization workflows.
Specifically, researchers are reaching a new state-of-the-art in NLP tasks using BERT embeddings. As for Elasticsearch, now you can index documents with sentence embeddings!
We thought this was a fantastic context to show others how we have found success building clinical search tools while we update our work to reflect the current state-of-the-art.
The Data
The dataset consists of ~44,000 research papers with nearly 29,000 of those articles about related COVID-19 and coronaviruses.
Our Approach
To make this rich information more accessible to expert researchers, we want to build a search engine. Since our last app, Elasticsearch has powerful new capabilities through the dense_vectors api to score document relevance based on cosine similarity between document and query embeddings.
BERT represents the state-of-the-art in many NLP tasks by introducing context-aware embeddings through the use of Transformers.
BERT embeddings work well on short text excerpts like paragraphs and so we form new Elasticsearch documents based on each paragraph. Then using a server hosting a BERT model, we extract embeddings for each paragraph and used the dense_vectors to index the thousands of research papers.

The Results
This repo includes everything needed to stand up a flask application running elasticsearch and BERT to index documents. This snappy app is able to return highly relevant content for a technical query.

Note the high quality results despite the lack of keyword matching to the query.